Research Interests
Technology and engineering
must be based on pure science;
the time for empirical
invention is long past
Kieth J. Laidler, "To Light Such
A Candle" Oxford University Press, 1998
The primary research interests in the Kelty group focus
on modeling and simulation of solid state surfaces and interfacial phenomena.
This includes application areas such as catalysis, polymer properties and
dynamics, biological materials including lipids, membrane
proteins and nucleic acids.
Some current research areas include:
-
Molecular
dynamics simulation of lipid membranes containing mixed dipalmitoylphosphatidylcholine (DPPC)
and dimyristoylphosphatidylcholine (DMPC) which differ only in the length of
the aliphatic tails by
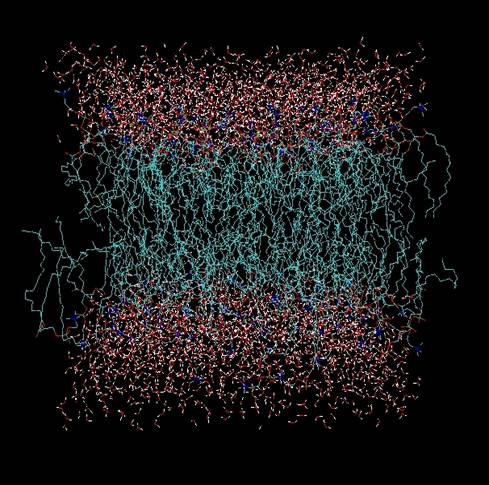 two methylene units. We have undertaken classical MD
simulations of DMPC/DHCP mixtures two determine inter-miscibility, diffusion
constants and structural properties of this
mixed-lipid system. The picture at left shows this system following 50
ns of simulation time. Future aspects of this project will include
addition of a membrane protein Bacteriorhodopsin, bR). The goal is to
determine how the protein's function is
influenced by the chemical composition of the lipid bilayer.
two methylene units. We have undertaken classical MD
simulations of DMPC/DHCP mixtures two determine inter-miscibility, diffusion
constants and structural properties of this
mixed-lipid system. The picture at left shows this system following 50
ns of simulation time. Future aspects of this project will include
addition of a membrane protein Bacteriorhodopsin, bR). The goal is to
determine how the protein's function is
influenced by the chemical composition of the lipid bilayer.
- One of the most powerful complementary techniques to
Scanning Tunneling Microscopy (STM) and Atomic Force Microscopy (AFM),
particularly at atomic resolution, is the ability to model image data based on a
hypothetical structure model. For example, we have been able to
correctly interpret STM images of the graphitic layer edge which show a
novel super-structure as resulting from a layer edge rearrangement of the
electronic structure and not to crystal reconstruction. We
have also investigated the local edge properties of Transition Metal Sulfides
(TMS) (NbS2) using both experimental STM and complementary EHTB and
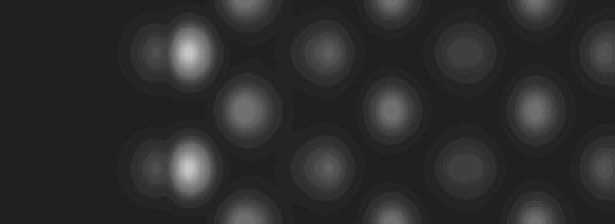
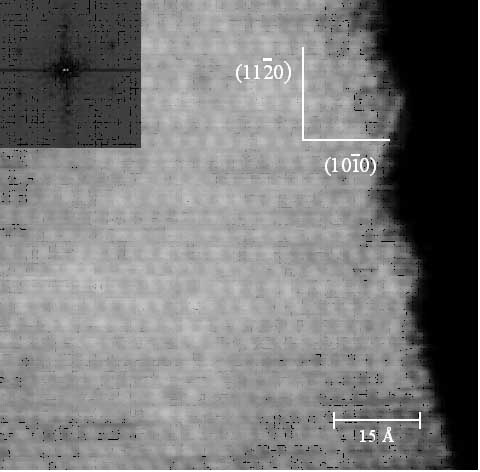
Density Functional Theory (DFT) computational methods. The image at
below left is experimental STM data of the edge region of NbS2 and
on the right id a simulated image of the edge.


- We are also investigating the equilibrium and
dynamical
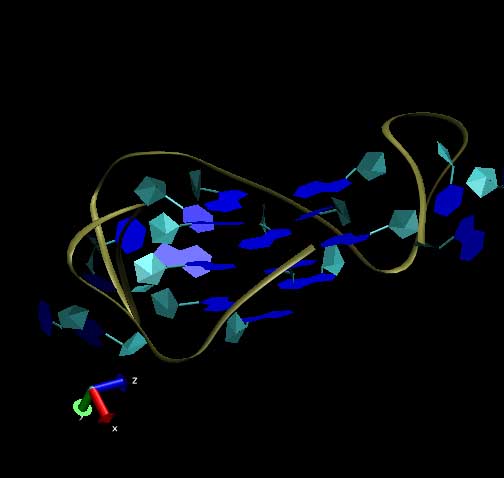 properties of guanine-rich quadruplex structures.
In normal duplex DNA, the two strands are bound together using
complementary base pairing (A-T and C-G). Quadruplexes are
composed of an unusual bonding arrangement in which four guanine residues
are bound to each other in each "step" of the ladder in the sequence.
We are currently investigating the oligomer (GGGTTA)4 in which three
quadruplexes are separated by the TTA linkages. The goal of this
project is to better understand the relative stability of alternative structure
types.
properties of guanine-rich quadruplex structures.
In normal duplex DNA, the two strands are bound together using
complementary base pairing (A-T and C-G). Quadruplexes are
composed of an unusual bonding arrangement in which four guanine residues
are bound to each other in each "step" of the ladder in the sequence.
We are currently investigating the oligomer (GGGTTA)4 in which three
quadruplexes are separated by the TTA linkages. The goal of this
project is to better understand the relative stability of alternative structure
types.
- Metal Oxides are also being investigated using
classical MD to investigate the properties of thin films of mixed metal oxides
as well as the surface properties of pure metal oxides.
 two methylene units. We have undertaken classical MD
simulations of DMPC/DHCP mixtures two determine inter-miscibility, diffusion
constants and structural properties of this
mixed-lipid system. The picture at left shows this system following 50
ns of simulation time. Future aspects of this project will include
addition of a membrane protein Bacteriorhodopsin, bR). The goal is to
determine how the protein's function is
influenced by the chemical composition of the lipid bilayer.
two methylene units. We have undertaken classical MD
simulations of DMPC/DHCP mixtures two determine inter-miscibility, diffusion
constants and structural properties of this
mixed-lipid system. The picture at left shows this system following 50
ns of simulation time. Future aspects of this project will include
addition of a membrane protein Bacteriorhodopsin, bR). The goal is to
determine how the protein's function is
influenced by the chemical composition of the lipid bilayer.

 properties of guanine-rich quadruplex structures.
In normal duplex DNA, the two strands are bound together using
complementary base pairing (A-T and C-G). Quadruplexes are
composed of an unusual bonding arrangement in which four guanine residues
are bound to each other in each "step" of the ladder in the sequence.
We are currently investigating the oligomer (GGGTTA)4 in which three
quadruplexes are separated by the TTA linkages. The goal of this
project is to better understand the relative stability of alternative structure
types.
properties of guanine-rich quadruplex structures.
In normal duplex DNA, the two strands are bound together using
complementary base pairing (A-T and C-G). Quadruplexes are
composed of an unusual bonding arrangement in which four guanine residues
are bound to each other in each "step" of the ladder in the sequence.
We are currently investigating the oligomer (GGGTTA)4 in which three
quadruplexes are separated by the TTA linkages. The goal of this
project is to better understand the relative stability of alternative structure
types.